The Stem Cell Commons
The number of high-throughput sequencing applications is ever-increasing which, along with consistently plummeting costs, has significantly increased the accessibility of this technology. However, progress has comparatively lagged behind in making bioinformatics solutions available to wet lab biologists that allow them to manage and explore their data as straightforwardly as they can now produce it.
Consequently, we have developed the Stem Cell Commons platform for the Harvard Stem Cell Institute to provide non-specialists with solutions for storing data, executing popular analysis workflows (e.g. RNA-seq, ChIP-seq), and visualizing results.
Our goal is that this platform encourages more stem cell biologists to pursue high-throughput sequencing experiments and particularly enables them to independently explore their data to foster new insights.
All data and tools are open source for the Stem Cell Commons are freely available and can be accessed through our public site:
https://hsci.stemcellcommons.org
For help, see The Stem Cell Commons Tutorial
Discover
Search for and
Discover
Over 200 Stem Cell-related Data Sets
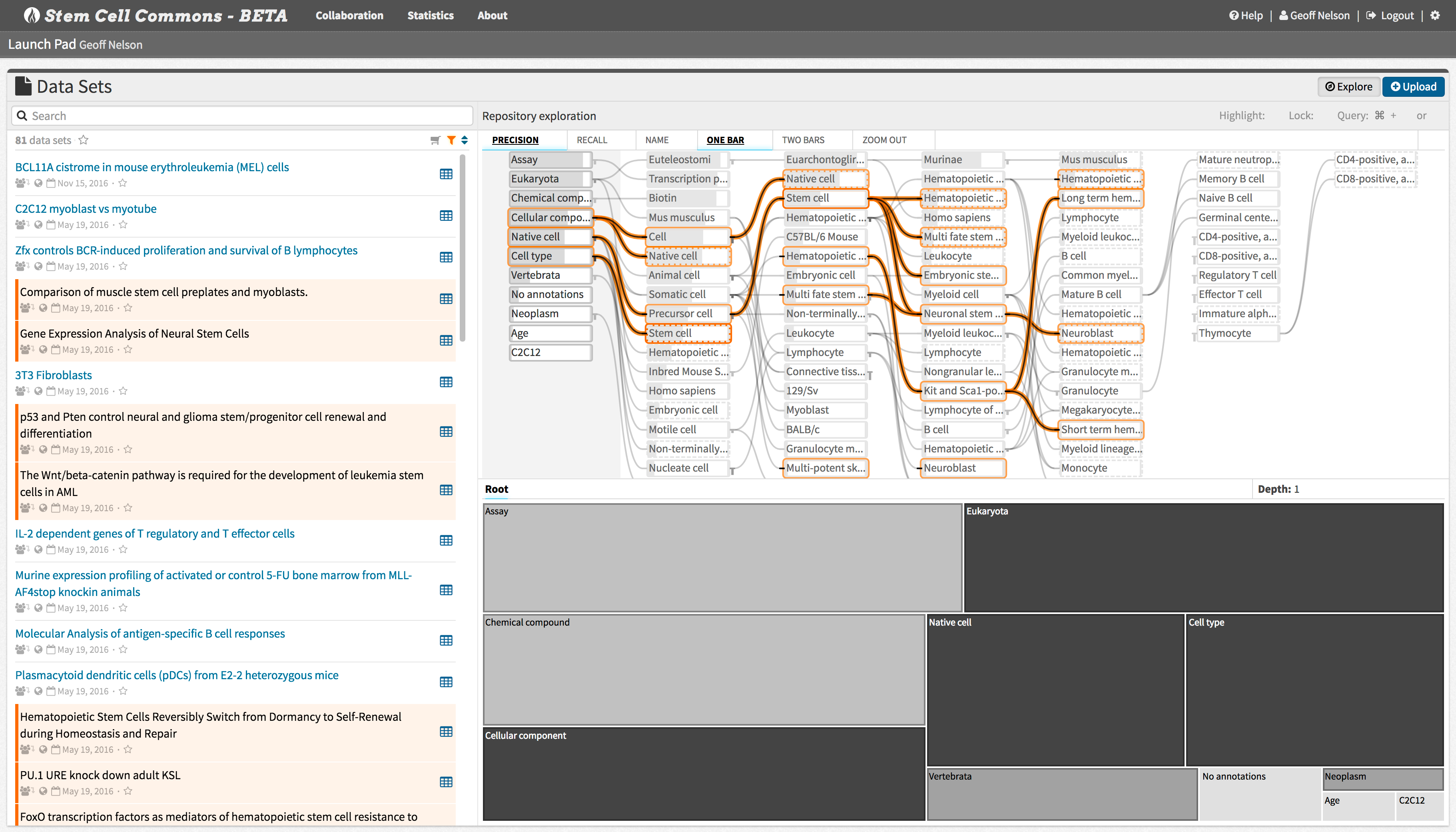
Find data relevant to your studies using full-text search and annotation-based visual exploration of stem cell-related transcriptomics and epigenomics data sets.
Manage
Upload and
Manage
Your Own Data Sets
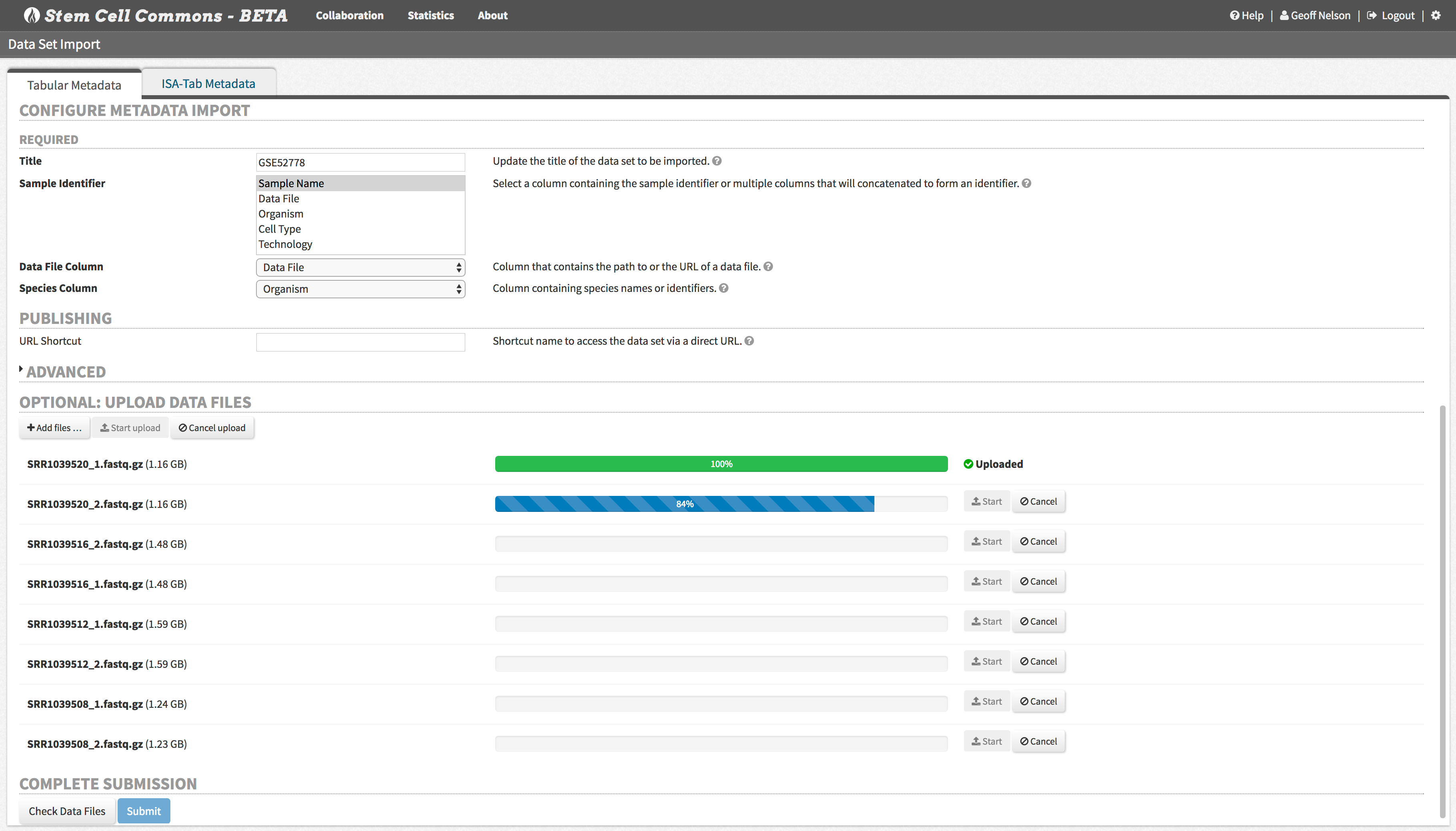
Import data files along with structured metadata to establish a searchable repository for all of your projects.
Analyze
Import and
Analyze
Your Own or Public Data Sets
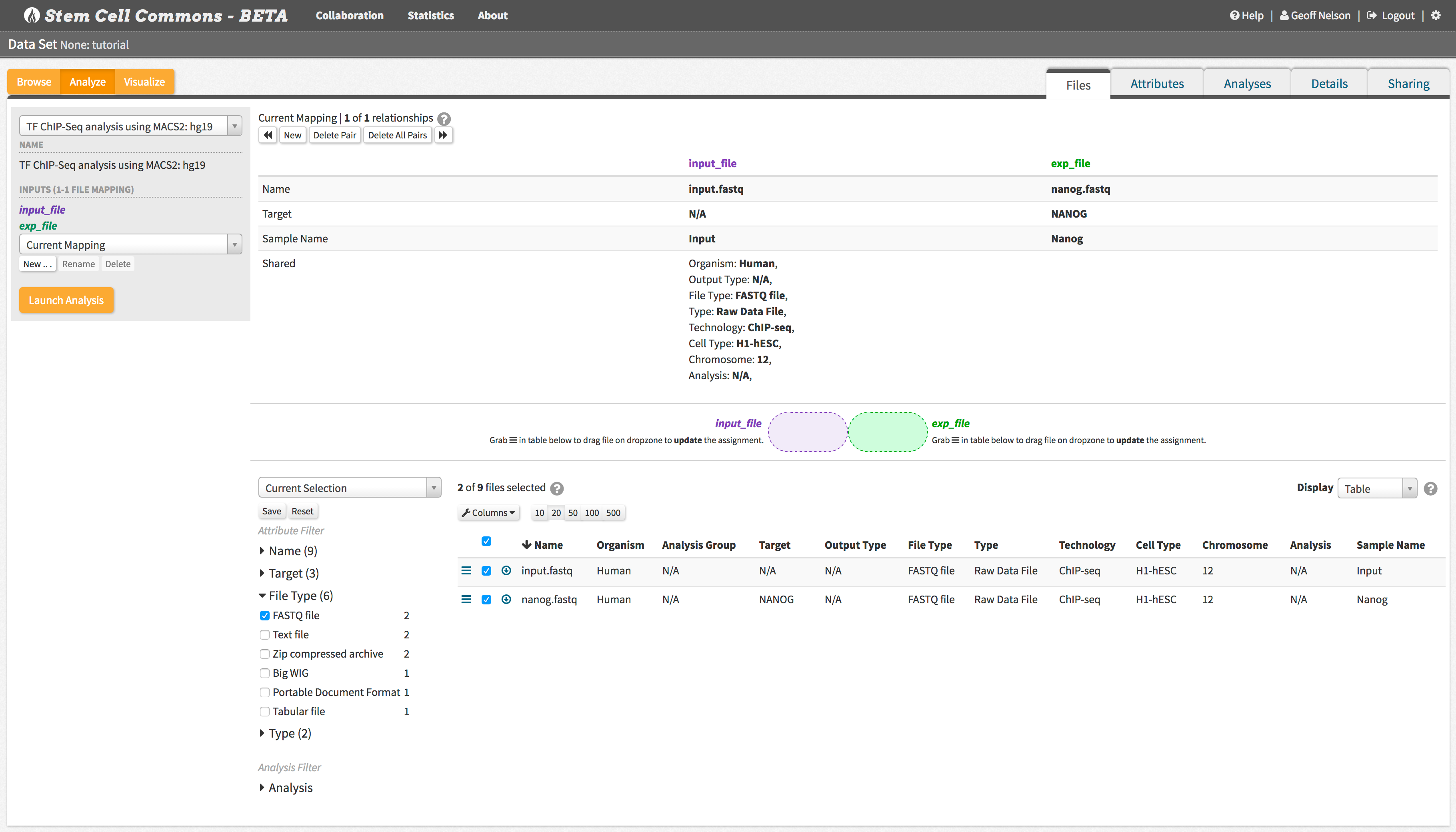
Launch workflows for FastQC, ChIP-seq peak calling, and RNA-seq quantification.
Collaborate
Invite and
Collaborate
With your Colleagues on your Analyses
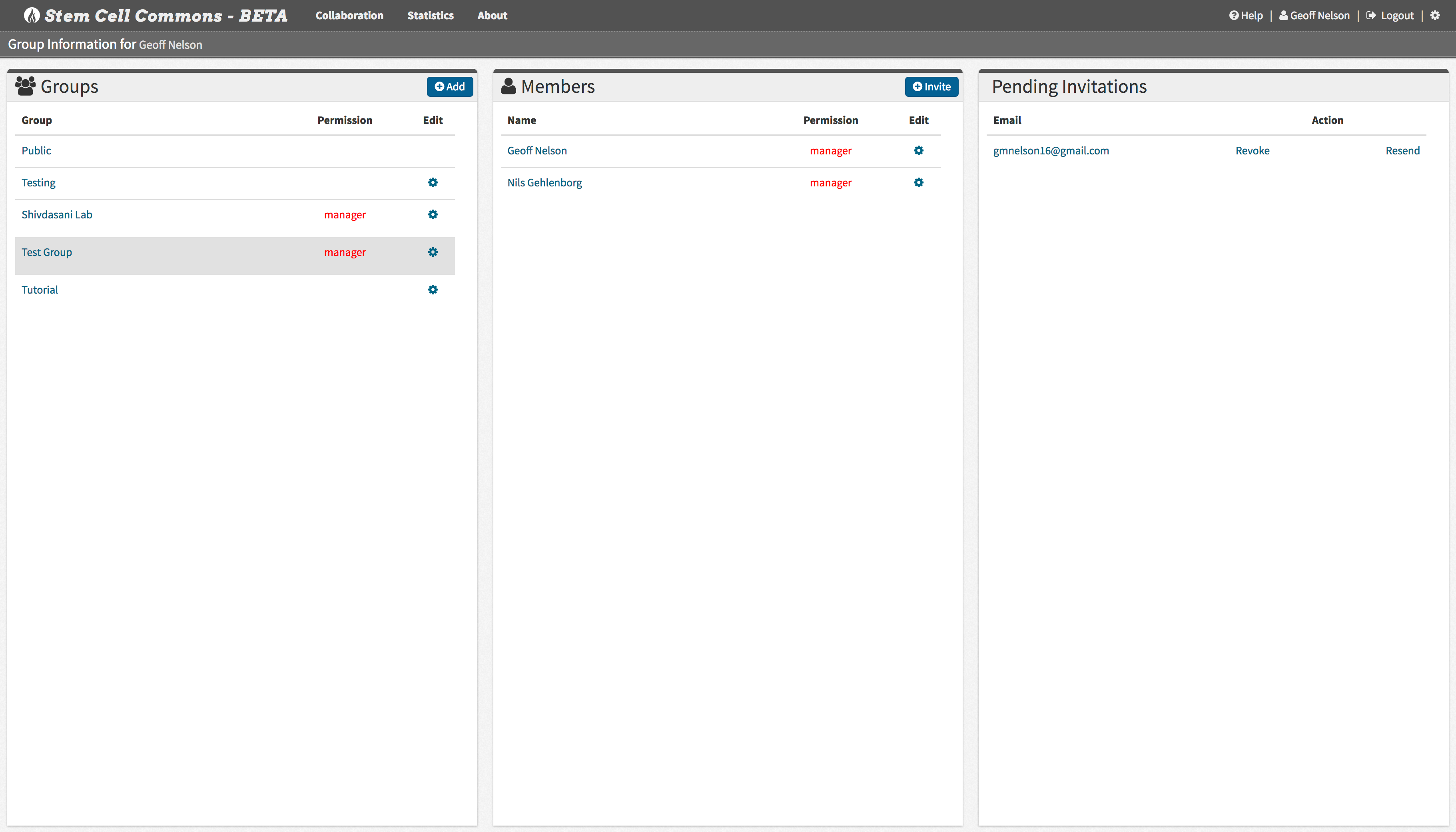
Add collaborators to your data sets/analyses by sending invitation emails directly through the Stem Cell Commons.
Reproduce
Access Tools that Allow You to
Reproduce
Studies and Document Findings
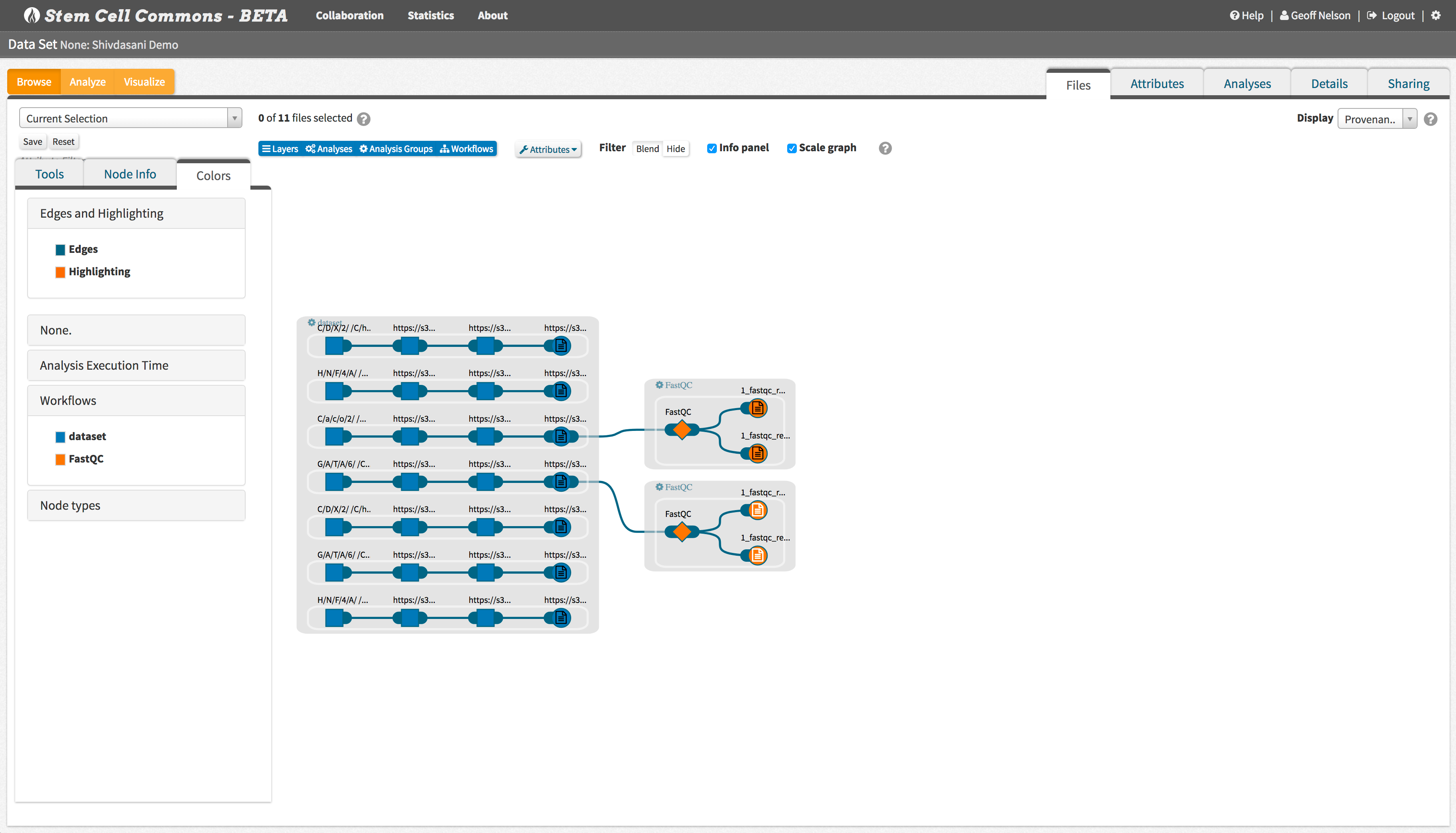
Use scalable visualization tools built into the Stem Cell Commons to review analyses and how files were generated.
Visualize
Explore and
Visualize
Results of your Analyses
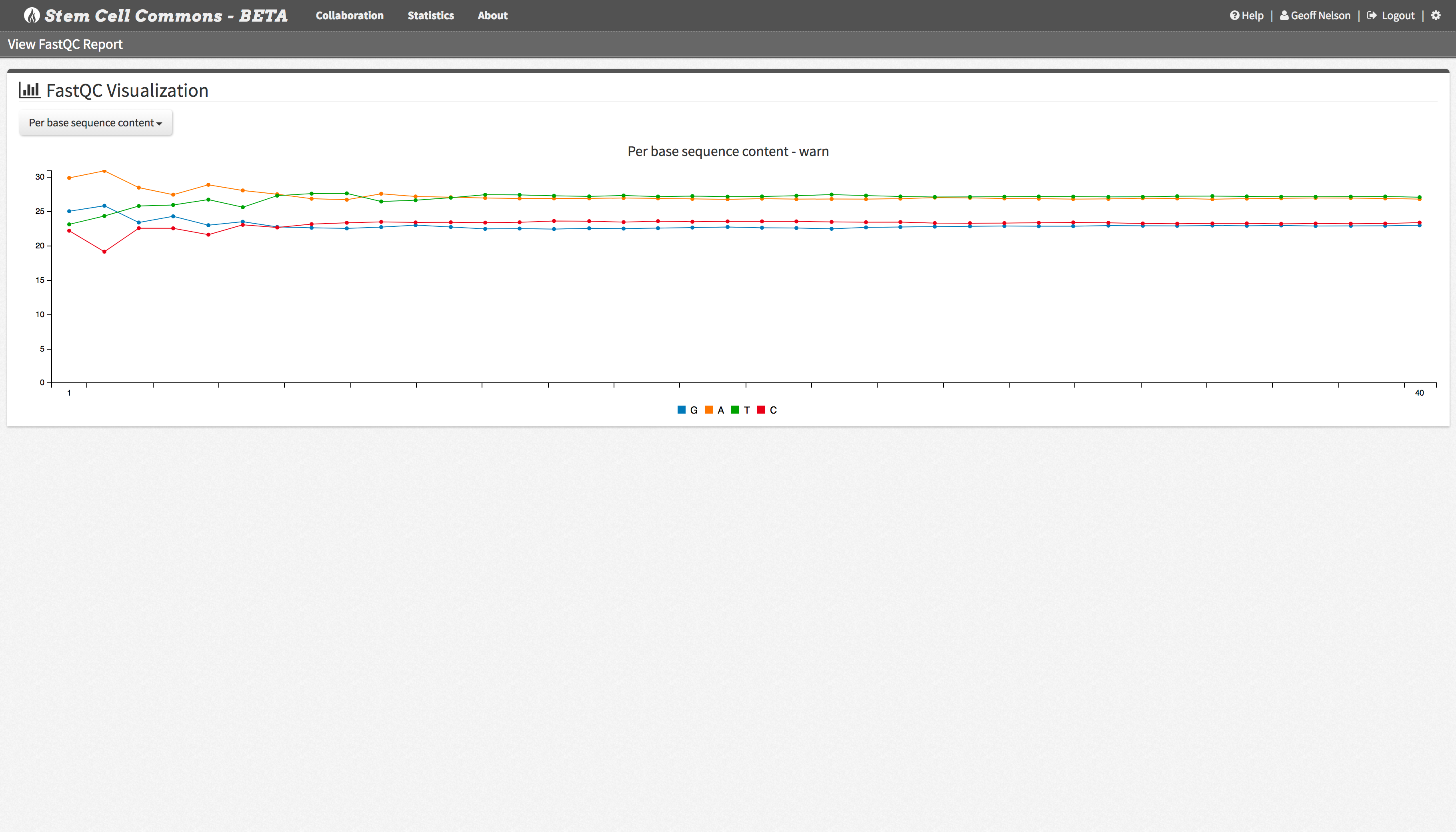
Launch the Integrative Genomics Viewer (IGV) directly from the Stem Cell Commons to explore genomic data or visualize FastQC results.
Publish
Share and
Publish
Data and Analysis Results
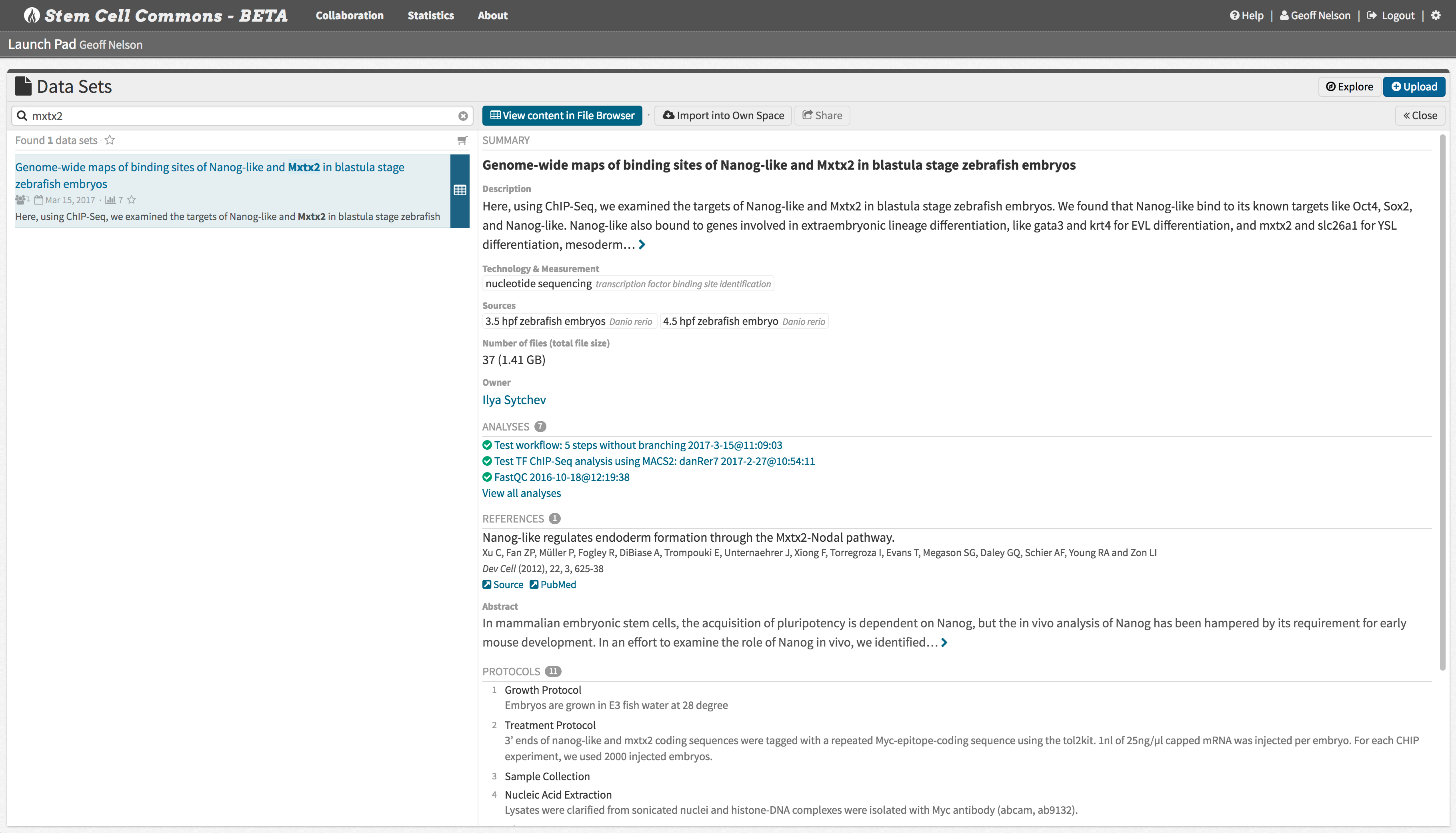
Data sets and associated analyses can be published at the click of a button to share your findings with the scientific community.
Team

Peter Park
Principal Investigator
Park Lab

Shannan Ho Sui
Co-investigator
Harvard Chan Bioinformatics Core

Nils Gehlenborg
Co-investigator
Gehlenborg Lab

Ilya Sytchev
Senior Bioinformatics Developer
Harvard Chan Bioinformatics Core

Jennifer Marx
Software Engineer
Gehlenborg Lab and Park Lab

Scott Ouellette
Software Developer
Gehlenborg Lab and Park Lab

Chuck McCallum
Software Developer
Gehlenborg Lab

Fritz Lekschas
Graduate student
Gehlenborg Lab
